Cell-free DNA (also known as circulating-free DNA) was discovered in human blood in 1948 (1), and it generally comprises fragments of nucleic acids that circulate in the blood without any cells attached to it (in other words, just DNA out in the wild). This did not catch on as a diagnostic modality until 1997, when Y chromosomal fetal DNA was extracted from the blood of pregnant women (2). These DNA fragments are released from the placenta into the maternal circulation, and this genetic fragment has been used to non-invasively screen women who are at high risk of trisomy 13, 18, and 21 (2). This modality has also been used to diagnose cancers (3, 4, 5) and solid-organ transplant rejection (6, 7), which circumvents the need for a tissue biopsy. Indeed, this modality has been called a “liquid biopsy.”
The idea behind this is as follows: cfDNA are derived from dying human cells and microorganisms that release their contents into the blood as they break down (1, 8). Compared to genomic DNA, cfDNA is much smaller, usually around 170bp. Due to this, healthy people have a concentration of cfDNA that ranges from 0-100ng/mL of plasma, sometimes exceeding 1500ng/mL (8). Human DNA accounts for anywhere from 90-99%, with microbial cfDNA making up a small fraction (0.08-4.85% for bacteria, 0-0.01% for fungi, 0.0-0.16% viruses, 9). This, of course, suggests that blood may not necessarily be sterile, but that is a topic for another day (for all intents and purposes, you should know that blood is normally sterile). Recently, the Karius test (10) has emerged as a new diagnostic modality that uses cfDNA to detect 1250 different bacteria, fungi, and viruses.
How does it work?
There are several advantages of using cfDNA for diagnostics. First, it is easy to “pick” it apart from the larger genomic DNA. Further, the half-life of cfDNA is actually quite short, around 10-15min (8), meaning that once the organism of interest is cured, you won’t have to worry about “dead DNA” floating around. Third, since it picks up cfDNA from anywhere in the circulation, you can get results from infections that are in difficult to culture areas, such as the CNS or other deep seated infection. Fourth, you are able to monitor the disease progression, as it is a quantitative method.
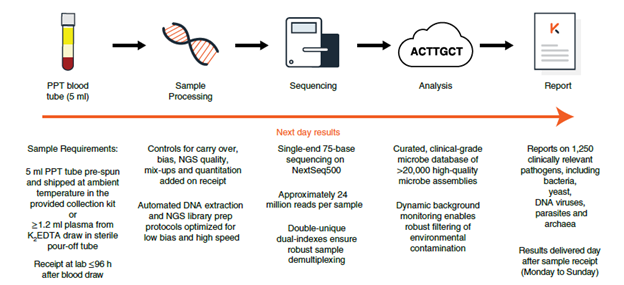
- At least 5cc of blood is drawn from the patient, preferably from a 21G or large bore needle to prevent hemolysis of cells
- This sample is then spun at low speed to separate plasma
- The sample is then shipped to the Karius facility overnight
- Once in the facility, cfDNA is extracted from the sample. There are several methods ,but ultimately they all do the same thing which is to get the sample you want from the rest of the plasma (11):
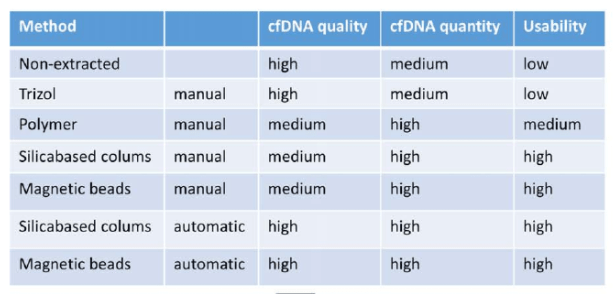
- From here, the sample is sequenced.
- The sequence is then analyzed against a negative control (remember that human DNA makes up to 99% of cfDNA and healthy patients may even have some translocation of microbial cfDNA that does not represent infection) in the database. Some microbes that are deemed to be contaminants will require a higher threshold of cfDNA prior to being flagged, compared to other more pathogenic organisms:
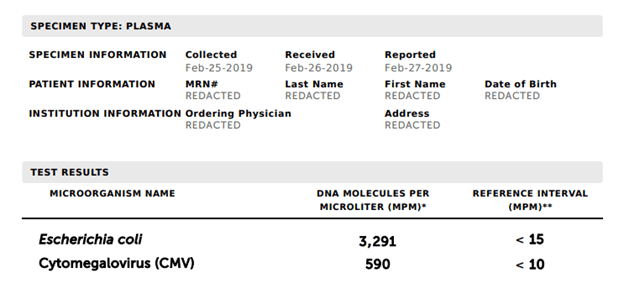
What is the data behind this?
A retrospective study of 10 immunocompromised patients found that cfDNA had 100% concordance 5 of the patients with a microbiological diagnosis and was able to find the etiology of 2 of the remaining 5 who did not have any culture positivity (12). A larger study of 79 pediatric patients found that the sensitivity/specificity of cfDNA was 92%/64% respectively compared to conventional methods, 77%/89% (13). In a case series of 9 patients with proven invasive fungal infections, 7/9 patients had concordant cfDNA test results, with one actually identifying an organism as Aspergillus lentulus, when conventional microbiology identified it as A. fumigatus complex (A. lentulus is morphologically identical to A. fumigatus, 14). The remaining 2 cases being negative were attributed to either delayed timing of plasma collection, and the other actually as below the threshold for detection. A review of the utility of the test on tuberculosis found the sensitivity in plasma ranged from 29-65% when compared to culture, and in urine it ranged from 43-79% (1):
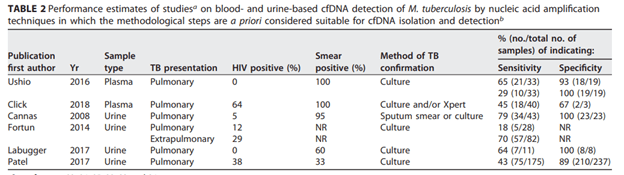
Notably here, it had a fairly high sensitivity for extrapulmonary tuberculosis, but this was only one study that was in the review. This assay has also been used in HIV patients with one case series describing its use in 4 patients (15).
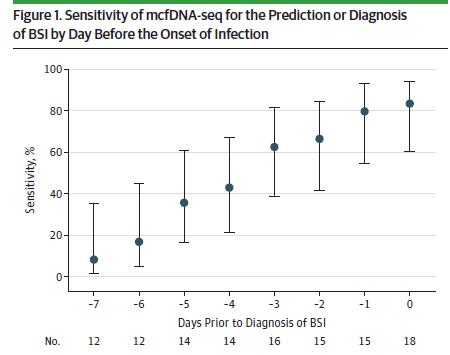
cfDNA may also be used for predicting infections in high risk individuals, before traditional cultures would have turned positive. In a retrospective study of 47 pediatric patients with hematological malignancies (16), blood samples from 3-7 days prior to infection was evaluated using cfDNA and compared to subsequent positive blood cultures. 19 episodes of BSI were found, of which 16 had predictive samples. The overall predictive sensitivity was 75%, (95% CI 51-90%) and specificity was 82% (95% CI 66-91%). These values were higher when taking out the one fungal infection (sensitivity 80%, specificity 91%):
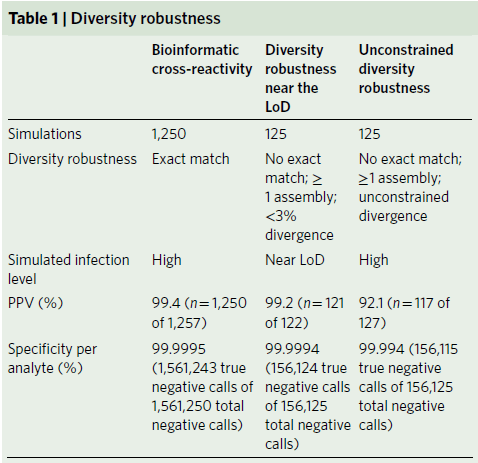
A study in Nature validated this modality (17). Using asymptomatic human plasma, 50 samples were produced to determine the specificity in context of environmental-contamination. Here, the overall specificity was 98%. Furthermore, cross-reactivity simulations showed a high specificity for organisms that may not necessarily be an exact match, though this goes down when the attempting to detect organisms that were not an exact match (higher divergence from database). The specificity, however, remained high here:
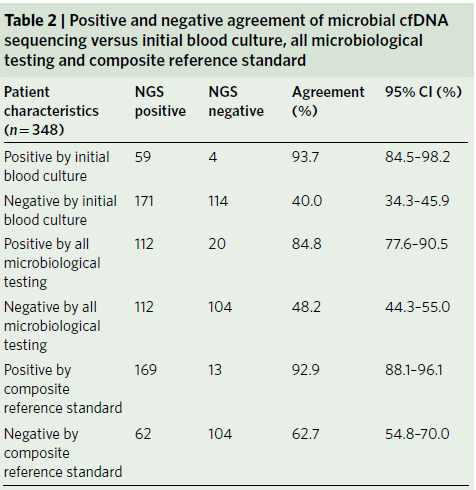
Clinically, the assay was tested in a cohort of 350 patients who presented to an emergency department meeting sepsis criteria. ¼ of these patients had received antibiotics in the week prior to presentation. Compared to blood cultures, cfDNA sequencing had a sensitivity of 93.7% (59/63 95% CI 84.5-98.2%). When comparing cfDNA to subsequent serology, NAT, tissue cultures, and blood cultures, the sensitivity dropped to 84.9% (112/132), leading to an overall sensitivity of 92.9% (169/182, 95% CI 88.1-96.1) with it being higher than all micro testing over seven days (132/348). In 166 samples without a microbiological diagnosis, cfDNA was detected in 104 (specificity 62.7%).
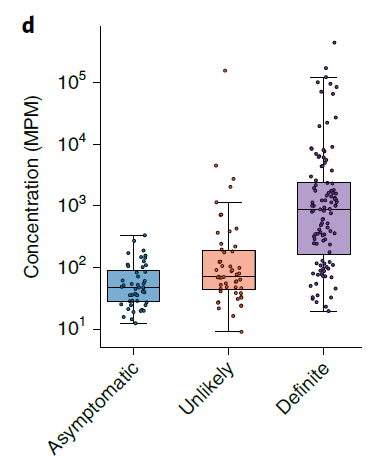
What about asymptomatic patients? 167 asymptomatic samples were tested, with 77.2% having no organisms identified, and the rest having significantly lower concentrations than those who were symptomatic:
cfDNA resulted in a shorter turn-around time when compared to conventional microbiology, 53h vs 92.4h (P-value 0.0004):
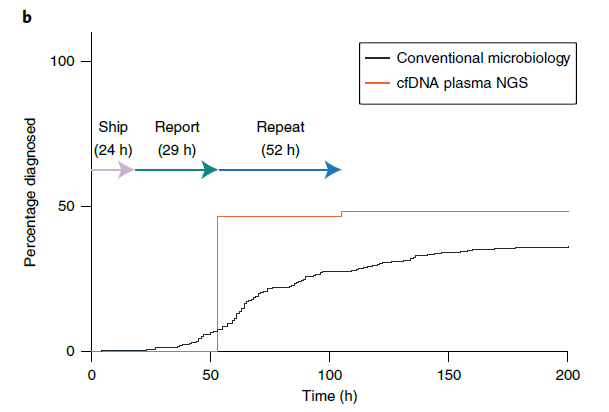
cfDNA seems to be an up and coming diagnostic modality for infectious disease, with its niche being its utility in culture-negative high-risk patients, and in those where biopsy may be prohibited due to its high risk nature. While it is out in use already, there is more data forthcoming, so we will continue to learn more about its utility in clinical practice.
References:
- Fernández-Carballo BL, Broger T, Wyss R, Banaei N, Denkinger CM. Toward the Development of a Circulating Free DNA-Based In Vitro Diagnostic Test for Infectious Diseases: a Review of Evidence for Tuberculosis. J Clin Microbiol. 2019 Mar 28;57(4):e01234-18. doi: 10.1128/JCM.01234-18. PMID: 30404942; PMCID: PMC6440766.
- Bianchi DW, Chiu RWK. Sequencing of Circulating Cell-free DNA during Pregnancy. N Engl J Med. 2018 Aug 2;379(5):464-473. doi: 10.1056/NEJMra1705345. PMID: 30067923.
- Fiala, C., Diamandis, E.P. New approaches for detecting cancer with circulating cell-free DNA. BMC Med 17, 159 (2019). https://doi.org/10.1186/s12916-019-1400-z
- Leal, A., van Grieken, N.C.T., Palsgrove, D.N. et al. White blood cell and cell-free DNA analyses for detection of residual disease in gastric cancer. Nat Commun 11, 525 (2020). https://doi.org/10.1038/s41467-020-14310-3
- Giacona MB, Ruben GC, Iczkowski KA, et al. Cell-free DNA in human blood plasma: length measurements in patients with pancreatic cancer and healthy controls. Pancreas. 1998 Jul;17(1):89-97. DOI: 10.1097/00006676-199807000-00012.
- Thongprayoon, C., Vaitla, P., Craici, I. M., Leeaphorn, N., Hansrivijit, P., Salim, S. A., Bathini, T., Rivera, F., & Cheungpasitporn, W. (2020). The Use of Donor-Derived Cell-Free DNA for Assessment of Allograft Rejection and Injury Status. Journal of clinical medicine, 9(5), 1480. https://doi.org/10.3390/jcm9051480
- Knight SR, Thorne A, Lo Faro ML. Donor-specific Cell-free DNA as a Biomarker in Solid Organ Transplantation. A Systematic Review. Transplantation. 2019 Feb;103(2):273-283. doi: 10.1097/TP.0000000000002482. PMID: 30308576.
- Han D, Li R, Shi J, Tan P, Zhang R, Li J. Liquid biopsy for infectious diseases: a focus on microbial cell-free DNA sequencing. Theranostics. 2020 Apr 7;10(12):5501-5513. doi: 10.7150/thno.45554. PMID: 32373224; PMCID: PMC7196304.
- Huang YF, Chen YJ, Fan TC, Chang NC, Chen YJ, Midha MK, Chen TH, Yang HH, Wang YT, Yu AL, Chiu KP. Analysis of microbial sequences in plasma cell-free DNA for early-onset breast cancer patients and healthy females. BMC Med Genomics. 2018 Feb 13;11(Suppl 1):16. doi: 10.1186/s12920-018-0329-y. PMID: 29504912; PMCID: PMC5836824.
- Blauwkamp, T.A., Thair, S., Rosen, M.J. et al. Analytical and clinical validation of a microbial cell-free DNA sequencing test for infectious disease. Nat Microbiol 4, 663–674 (2019). https://doi.org/10.1038/s41564-018-0349-6
- Volckmar AL, Sültmann H, Riediger A, Fioretos T, Schirmacher P, Endris V, Stenzinger A, Dietz S. A field guide for cancer diagnostics using cell-free DNA: From principles to practice and clinical applications. Genes Chromosomes Cancer. 2018 Mar;57(3):123-139. doi: 10.1002/gcc.22517. Epub 2017 Dec 20. PMID: 29205637.
- Camargo JF, Ahmed AA, Lindner MS et al. Next-generation sequencing of microbial cell-free DNA for rapid noninvasive diagnosis of infectious diseases in immunocompromised hosts [version 1; peer review: 1 approved]. F1000Research 2019, 8:1194 (https://doi.org/10.12688/f1000research.19766.1)
- Rossoff J, Chaudhury S, Soneji M, Patel SJ, Kwon S, Armstrong A, Muller WJ. Noninvasive Diagnosis of Infection Using Plasma Next-Generation Sequencing: A Single-Center Experience. Open Forum Infect Dis. 2019 Aug 1;6(8):ofz327. doi: 10.1093/ofid/ofz327. PMID: 31375834; PMCID: PMC6677669.
- Hong DK, Blauwkamp TA, Kertesz M, Bercovici S, Truong C, Banaei N. Liquid biopsy for infectious diseases: sequencing of cell-free plasma to detect pathogen DNA in patients with invasive fungal disease. Diagn Microbiol Infect Dis. 2018 Nov;92(3):210-213. doi: 10.1016/j.diagmicrobio.2018.06.009. Epub 2018 Jun 22. PMID: 30017314.
- Zhou, Yang & Hemmige, Vagish & Dalai, Sudeb & Hong, David & Muldrew, Kenneth & Al Mohajer, Mayar. (2019). Utility of Whole-Genome Next-Generation Sequencing of Plasma in Identifying Opportunistic Infections in HIV/AIDS. The Open AIDS Journal. 13. 7-11. 10.2174/1874613601913010007.
- Goggin KP, Gonzalez-Pena V, Inaba Y, Allison KJ, Hong DK, Ahmed AA, Hollemon D, Natarajan S, Mahmud O, Kuenzinger W, Youssef S, Brenner A, Maron G, Choi J, Rubnitz JE, Sun Y, Tang L, Wolf J, Gawad C. Evaluation of Plasma Microbial Cell-Free DNA Sequencing to Predict Bloodstream Infection in Pediatric Patients With Relapsed or Refractory Cancer. JAMA Oncol. 2020 Apr 1;6(4):552-556. doi: 10.1001/jamaoncol.2019.4120. Erratum in: JAMA Oncol. 2020 Feb 27;: PMID: 31855231; PMCID: PMC6990667.
- Blauwkamp TA, Thair S, Rosen MJ, Blair L, Lindner MS, Vilfan ID, Kawli T, Christians FC, Venkatasubrahmanyam S, Wall GD, Cheung A, Rogers ZN, Meshulam-Simon G, Huijse L, Balakrishnan S, Quinn JV, Hollemon D, Hong DK, Vaughn ML, Kertesz M, Bercovici S, Wilber JC, Yang S. Analytical and clinical validation of a microbial cell-free DNA sequencing test for infectious disease. Nat Microbiol. 2019 Apr;4(4):663-674. doi: 10.1038/s41564-018-0349-6. Epub 2019 Feb 11. PMID: 30742071.

1 comments on “Curious About Karius”