This is a topic that can be confusing, given the different numbers of assays and treatment algorithms that exist. Different hospitals have their own algorithm so this makes this a bit confusing when going from one shop to the next. In general, testing (at least nowadays) tends to fall into 2 general categories:
- Enzyme immunoassay (i.e. lateral flow, ELISA) – these target either the actual toxin products (A/B) or a preserved enzyme (glutamate dehydrogenase aka GHD)
- Nucleic acid amplification – targeting specifically toxin producing genes
These are “newer” testing modalities. Prior to this, we used very cumbersome methods for diagnosis that were labor intensive and time consuming.
- CCCNA (Cell Culture Cytotoxicity Neutralization Assay) – you put in some of the sample stool and put in on top of a cell line. Let it incubate for 24-48hrs and see if there is toxin-induced cytopathic effects. IF this is the case, add in either C.diff or C.sordellii antiserum and if it stops -> diagnosis.
- Of course this requires 1-2 days of incubation, plus the sensitivity ranges from 65% to 90%. It is used as a gold-standard in some reference labs, but honestly, this kind of is not a convenient test (again, takes like 2 days!)
- Toxigenic culture – most of the studies on new assays now use this as their reference. Unfortunately, there are many different ways of doing this that it is difficult to standardize. In essence, you put stool in either an anaerobic agar or broth culture that selects for C.diff, usually a cycloserine, cefoxitin, and fructose agar (CCFA). How the stool is treated prior to inoculation depends, either using heat shock or alcohol shock to kill off everything but the damn spores. From here, you let it culture for at least 48hrs and up to 7 days, then:
- Identify organism based on gram stain, colony, morphology, odor (aka horse barn odor, whatever that is), biochemical testing, or MALDI-TOF
- THEN, you need to test to see if it makes toxin, either CCCNA above OR a toxin EIAs (see below).
- No one uses this, thankfully
What do the US guidelines state as being the test of choice. Well they’re not really helpful here:

Why is this? If patients have no symptoms of diarrhea or C.diff colitis, they can still be colonized with a strain that is able to make toxin and not have disease. The NAAT picks up the genes (i.e. it is more sensitive) but the toxin test actually picks up the toxin (i.e. more specific). From the guidelines:
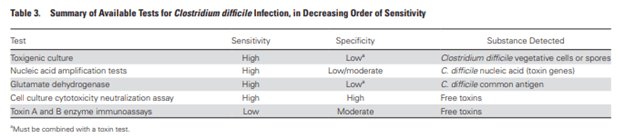
Let’s take a look at how each of theses assays peforms
- Glutamate Dehydrogenase and Toxin A/B EIAs
Lets review a bit of EIAs

The “target specimen” that is used for C.diff include glutamate dehydrogenase, which is widely distributed in nature, and the clostridial toxins, A and B. Of course, there are a variety of EIAs, just to make things more difficult. There is the ELISA, CIMA, and lateral flow assay:
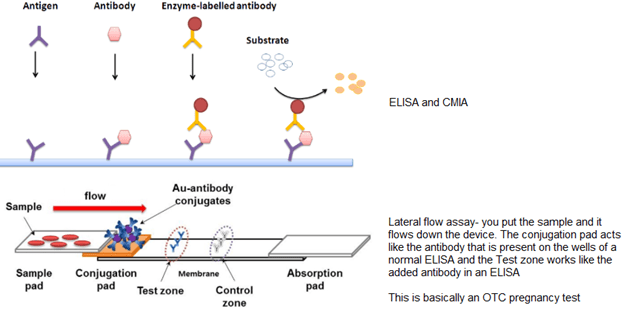
A few things to talk about each “target:”
- GDH – hit is a metabolic enzyme that is expressed in all C.diff isolates (Toxigenic and non-toxigenic) and the assay cross-reacts with C. Sordellii. In general, GDH tends to have a higher sensitivity than toxin A/B, because almost all strains have this
- Toxin A/B – this is more specific for that GDH, but less sensitive. The reason for the varying sensitivity has been cited to be due to circulating strain types (with differences of toxin expression(, geography, and the “gold standard” assay use for comparison.
How do each of these perform?
A study from China (1) evaluated the use of GDH as a screening method in 416 stool samples. This was compared to toxigenic culture and toxin A/B EIA. Notably, the specificity of both were fairly high but GDH performed much better as a screening modality:

In a retrospective study (2) evaluating the (at the time) new GDH/toxin EIA combo, both GDH and PCR ended up doing very well in terms of sensitivity when compared to toxin EIA (note, this was using the at the time new GDH/toxin EIA; PCR was used as a tie breaker if there were discrepant results such as +GDH/- toxin or vice versa).
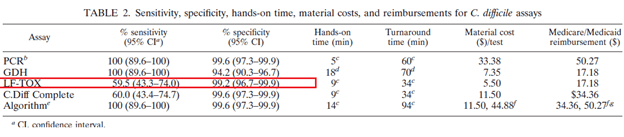
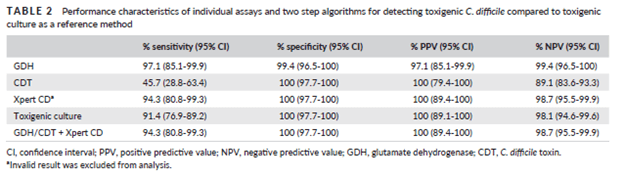
In another study comparing GDH/toxin EIA and the Cepheid Xpert for C.diff (3, 432 samples were analyzed. The standard was cell culture cytoxicity neutralization assay. They compared the sensitivity and specificity across each test individually and algorithms. As it can be seen, Xpert had the highest sensitivity and specificity when compared to other tests:

Interestingly, GDH + Xpert had a lower sensitivity and GDH + EIA actually had a lower sensitivity when compared to EIA alone. This suggest the utility in GDH as a potential screening tool. Indeed, a review (4) and meta-analysis (5) suggests that GDH is highly sensitive, a finding that is highlighted in the C.diff guidelines (6).

In said meta-analysis (5) the use of GDH, the pooled sensitivity was 91.1% (95% CI 87.1-94%) while specificity was 91.2% (95% CI 89.2-92.8%).
One thing to keep in mind is the heterogeneity of these studies. In 31 studies, CCNA was used as reference while toxin culture was used in 23. The assay evaluated also varied by study, but overall seems GDH had a fairly good sensitivity overall:

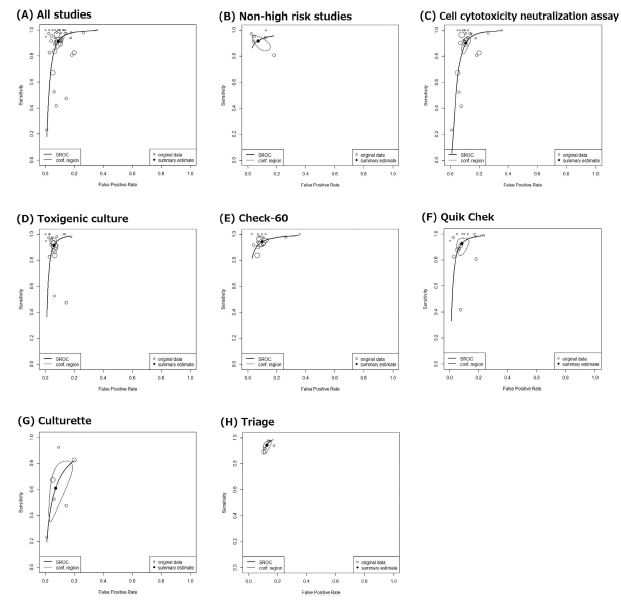
Notably, toxin EIA does not seem to do well as a screening tool. Indeed, in a retrospective (7) found the sensitivity of EIA compared to toxigenic culture was about 47%:
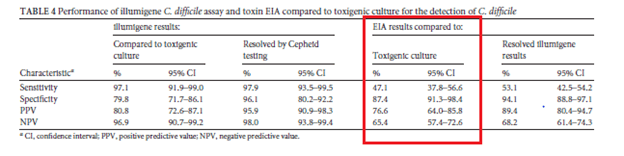
Further data comes from 2 additional studies (8. 9) one comparing the Gene Ohm Cdiff PCR with a GDH/toxigenic culture and toxin EIA as part of an algorithm and another one comparing it to CCNA direct toxin B showed comparable or even better sensitivity:
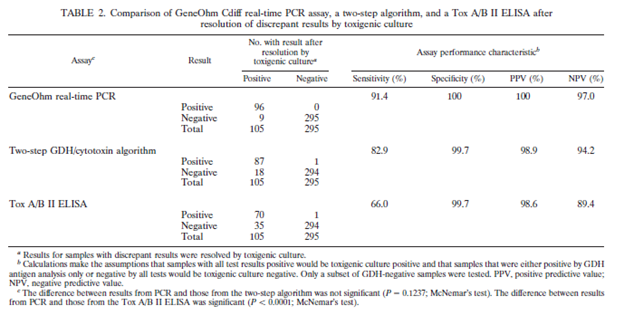
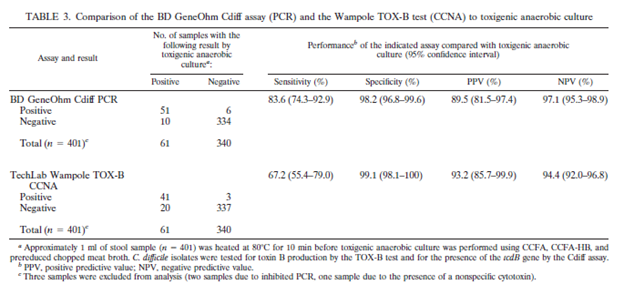
The biggest review on the topic (4) did evaluate several toxin EIA assays and found a huge variability in the sensitivity of each assay, varying from 42.5% up to 98.7%:
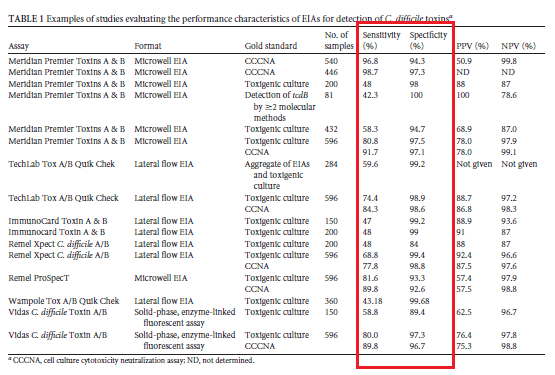
In summary, GDH tends to have a much higher sensitivity than toxin EIA, but toxin EIA has a higher specificity. So if your hospital uses a GDH assay, then a reasonable algorithm is as follows:

What is the clinical significance of a discrepant result in a GDH/toxin EIA?
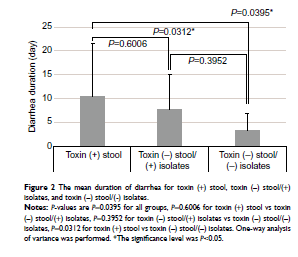
In a retrospective study (14), 68 stool specimens who were GDH positive were tested for toxin presence. In those, 29 ended up being negative. These specimens were then put into culture to increase the toxin yield. This resulted in 14 of the original toxin negative samples resulting in being toxin positive. Despite this, those who were originally toxin negative and ultimately tested toxin positive, they may not clinically behave differently from those who were toxin negative on isolate:
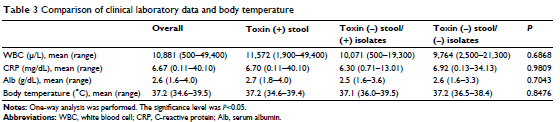

You may notice that NAAT is only used if test is equivocal in this algorithm. This is mostly due to the fact NAAT tend to be more expensive. Having said that:
- NAAT
Data for nucleic acid amplification testing tends to be favorable for it. For instance, in one Korean study (11), 199 stool samples using either GDH in conduction with C. diff toxin assay as well as with Gene Xpert. Not surprisingly, C diff toxin assay had the lowest sensitivity, but seems that GDH/CDT lateral flow assay did not provide additional advantage to the Gene Xpert assay when used by itself.
The studies cited above (8, 9) also suggest the sensitivity of PCR tends to be higher than that of toxin A/B but in some cases it may not be as sensitive as GDH, but this depends on the assay. In a meta-analysis (12) of 72 studies pooling several types of assays, they found the sensitivity for NAAT by itself was 95% (95% CI 94-96%) whil specificity was 98% (95% CI 97-98%). This is more sensitive than GDH -> NAAT strategy or even GDH-> Toxin ->NAAT strategy, though the specificity increases as you add more tests:

Further, they also pooled the sensitivity and specificity of all types of NAAT, with the Gene Xpert being the most sensitive and still fairly specific.

This was also found in the review (4) cited previously:

Having said that, it sounds like the sensitivity may depend on the specimen is collected and treated. For instance, in a study from Kuwait (13) the sensitivity of Gene Xpert was roughly 76.9%, which was better than that of a C.diff Quick Check Complete assay (an EIA for GDH and toxin). When the samples were inoculated in cooked meat broth, the sensitivity for both increased to 100%:
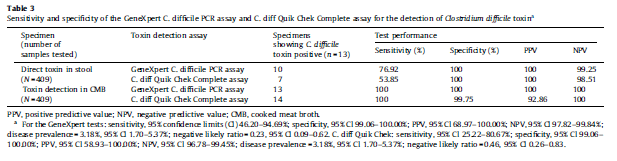
It should be mentioned the samples were from the community, rather than in-patient, however how this one study plays in to the other literature is difficult to say.
While there are several molecular assays, the major ones used in clinical practice are Cepheid GeneXpert and Nanosphere Verigene. Both detect tcdB (for toxin B), nucleotide 117 deletion on tcdC as a surrogate for 027/NAP1/Bl strain (aka the hypervirulent strain), and tcdA (for toxin A).
What is the caveat to NAAT? The biggest one is that it may pick up colonization, rather than true infection which can be difficult to discern. This is why the IDSA guidelines are framed as such:
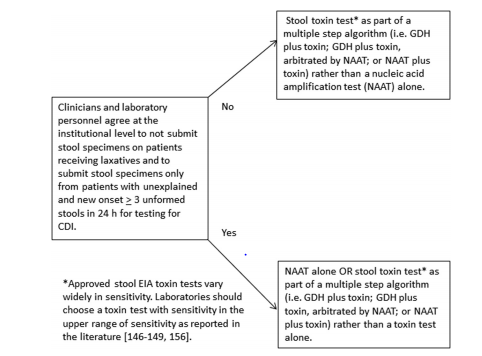
If you have a clinical syndrome suggestive of C.diff colitis, it is reasonable to go and proceed with a sensitive and relatively specific test with quick turnaround. This is why some labs turn down formed specimens. When someone has no diarrhea but are colonized with toxigenic C.diff, do you treat? If the lab makes no recommendation as to turn down formed stools, then the GDH followed by a toxin antigen is a reasonable approach, since you need the toxin to have disease.
Here is a reasonable approach to diagnosis:
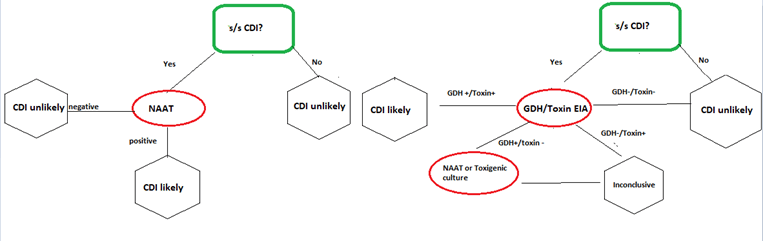
If you are ever in a situation where a NAAT was sent off and turned positive where the suspicion of C.diff is low (i.e. formed stool), then sending a toxigenic EIA is reasonable.
TL;DR
- The most sensitive testing modalities include EIA for GDH and Nucleic acid amplification
- Toxin EIA is very specific, but not very sensitive
- If lab only takes loose stools, then NAAT is a reasonable first-choice test without subsequent testing needed
- If stool is formed or if NAAT is not available, then GDH followed by either NAAT OR toxin EIA is reasonable as well
References:
- Cheng JW, Xiao M, Kudinha T, et al. The Role of Glutamate Dehydrogenase (GDH) Testing Assay in the Diagnosis of Clostridium difficile Infections: A High Sensitive Screening Test and an Essential Step in the Proposed Laboratory Diagnosis Workflow for Developing Countries like China. PLoS One. 2015;10(12):e0144604. Published 2015 Dec 11. doi:10.1371/journal.pone.0144604
- Sharp, Susan E et al. “Evaluation of the C.Diff Quik Chek Complete Assay, a new glutamate dehydrogenase and A/B toxin combination lateral flow assay for use in rapid, simple diagnosis of clostridium difficile disease.” Journal of clinical microbiology vol. 48,6 (2010): 2082-6. doi:10.1128/JCM.00129-10
- Novak-Weekley, Susan M., et al. “Clostridium Difficile Testing in the Clinical Laboratory By Use of Multiple Testing Algorithms.” Journal of Clinical Microbiology, vol. 48, no. 3, 2010, pp. 889-93.
- Burnham CA, Carroll KC. Diagnosis of Clostridium difficile infection: an ongoing conundrum for clinicians and for clinical laboratories. Clin Microbiol Rev. 2013;26(3):604-630. doi:10.1128/CMR.00016-13
- Arimoto J, Horita N, Kato S, et al. Diagnostic test accuracy of glutamate dehydrogenase for Clostridium difficile: Systematic review and meta-analysis. Sci Rep. 2016;6:29754. Published 2016 Jul 15. doi:10.1038/srep29754
- L Clifford McDonald, Dale N Gerding, Stuart Johnson, Johan S Bakken, Karen C Carroll, Susan E Coffin, Erik R Dubberke, Kevin W Garey, Carolyn V Gould, Ciaran Kelly, Vivian Loo, Julia Shaklee Sammons, Thomas J Sandora, Mark H Wilcox, Clinical Practice Guidelines for Clostridium difficile Infection in Adults and Children: 2017 Update by the Infectious Diseases Society of America (IDSA) and Society for Healthcare Epidemiology of America (SHEA), Clinical Infectious Diseases, Volume 66, Issue 7, 1 April 2018, Pages e1–e48, https://doi.org/10.1093/cid/cix1085
- Humphries RM, Uslan DZ, Rubin Z. Performance of Clostridium difficile toxin enzyme immunoassay and nucleic acid amplification tests stratified by patient disease severity. J Clin Microbiol. 2013;51(3):869-873. doi:10.1128/JCM.02970-12
- Stamper PD, Alcabasa R, Aird D, et al. Comparison of a commercial real-time PCR assay for tcdB detection to a cell culture cytotoxicity assay and toxigenic culture for direct detection of toxin-producing Clostridium difficile in clinical samples. J Clin Microbiol. 2009;47(2):373-378. doi:10.1128/JCM.01613-08
- Kvach EJ, Ferguson D, Riska PF, Landry ML. Comparison of BD GeneOhm Cdiff real-time PCR assay with a two-step algorithm and a toxin A/B enzyme-linked immunosorbent assay for diagnosis of toxigenic Clostridium difficile infection. J Clin Microbiol. 2010;48(1):109-114. doi:10.1128/JCM.01630-09
- Gateau C, Couturier J, Coia J, Barbut F. How to: diagnose infection caused by Clostridium difficile. Clin Microbiol Infect. 2018;24(5):463-468. doi:10.1016/j.cmi.2017.12.005
- Seo, Ja Young, et al. “Laboratory Diagnosis of Clostridium Difficile Infection: Comparison of Techlab C. Diff Quik Chek Complete, Xpert C. Difficile, and Multistep Algorithmic Approach.” Journal of Clinical Laboratory Analysis, vol. 31, no. 6, 2017.
- Kraft CS, Parrott JS, Cornish NE, et al. A Laboratory Medicine Best Practices Systematic Review and Meta-analysis of Nucleic Acid Amplification Tests (NAATs) and Algorithms Including NAATs for the Diagnosis of Clostridioides (Clostridium) difficile in Adults [published correction appears in Clin Microbiol Rev. 2019 Aug 28;32(4):]. Clin Microbiol Rev. 2019;32(3):e00032-18. Published 2019 May 29. doi:10.1128/CMR.00032-18
- Jamal W, Pauline EM, Rotimi VO. Comparative performance of the GeneXpert C. difficile PCR assay and C. diff Quik Chek Complete kit assay for detection of Clostridium difficile antigen and toxins in symptomatic community-onset infections. Int J Infect Dis. 2014;29:244-248. doi:10.1016/j.ijid.2014.10.025
- Yuhashi K, Yagihara Y, Misawa Y, et al. Diagnosing Clostridium difficile-associated diarrhea using enzyme immunoassay: the clinical significance of toxin negativity in glutamate dehydrogenase-positive patients. Infect Drug Resist. 2016;9:93-99. Published 2016 May 27. doi:10.2147/IDR.S105429
